Get the case study as a PDF.
TCR-Accepting T-Cell Platform Enables T-Cell Engineering with Enhanced Specificity and Prediction of Cross-Reactivity
Background
T-cell receptor (TCR) engineering is an appealing tool for cancer immunotherapy and is based on recognizing tumor antigens presented by major histocompatibility complex (MHC) molecules. Despite its significant promise, the engineering of TCR is complicated by a poor correlation between TCR affinity and functional response that can lead to cross-reactivity.
This study reports the development of a TCR-accepting T-cell (TnT) platform for the functional display, engineering, and cross-reactivity profiling of tumor-specific TCRs.
Study Description
Goal: validate the application of the TnT platform for TCR engineering and cross-reactivity profiling.
TNT platform was developed using the JurkatE6-1 cells subjected to CRISPR-Cas9 gene editing.
Using the platform, one of the reconstituted TCRs, MAGE-A3 TCRA3, was selected and validated by FACS based on CD3 surface expression, HLA-A*0101 EVDPIGHLY-specific Dextramer® binding, and antigen-induced signaling. Deep sequencing of CDR3β was done to identify TCR variants enriched across selection steps. To reveal variants with enhanced recognition of MAGE-A3, combinatorial TCR mutagenesis screening was performed, followed by functional cross-reactivity profiling.
Results
- Deep mutational scanning of CDR3β revealed enrichment of wild-type MAGE-A3 TCRA3 sequences in most selections, except for antigen-induced signaling, in which wild type TCRA3 was depleted (Fig.1A, SEL 2B boxes)
- MAGE-A3 TCRA3 reconstitution in TnT cells was confirmed by EVDPIGHLY-specific MHC I Dextramer® staining by FACS (Fig. 1B)
- Among selected TCRA3 mutants with the enrichment above wild-type TCRA3, S3G, P4L, M6L, and A7V displayed the largest increases in terms of binding and signaling (Fig. 1C).
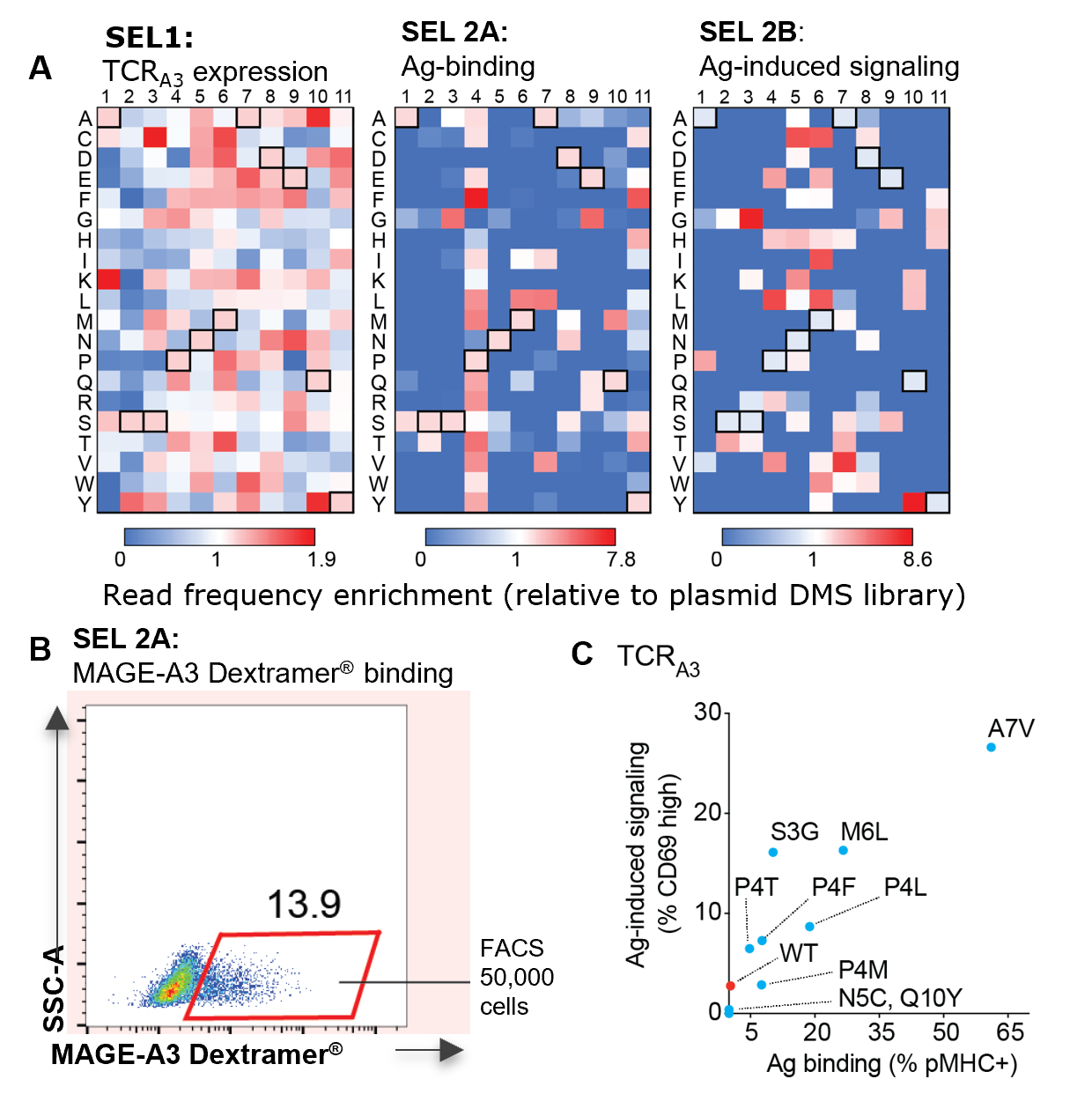
Fig.1. A. Heatmaps for the enrichment of sequencing reads from TCRA3 variants relative to their occurrence in the starting plasmid DMS libraries. Outlined boxes show wild-type CDR3β residues. B. TnTcells were transfected with CDR3β gRNA and HDR templates encoding the TCRA3 DMS library. Example of FACS selection based on binding to EVDPIGHLY-specific MHC I Dextramer®. C. Graph displays the levels of antigen binding and antigen-induced signaling in TCRA3 single mutants relative to wild-type TCRA3.
Conclusions
- The TnT platform is a valuable technology for the engineering of TCRs with enhanced functional and safety profiles
- Targeted TCR reconstitution for the generation of TCR-T cells can be assessed by detecting binding to cognate peptide-MHC I Dextramer® by flow cytometry.

